Page 55 - Haematologica Vol. 107 - September 2022
P. 55
ARTICLE - Genetic abnormalities in older ALL patients (0/56), JAK2 (0/53) or ABL2 (0/52) rearrangements were
detected.
Copy number alterations
SNP arrays were performed on diagnostic bone marrow samples from 78 of the 210 patients (49 from UKALL14 and 29 from UKALL60+) using the Illumina CytoSNP 850k (n=51) and Affymetrix Cytoscan HD (n=27) arrays. The SNP array cohort was reasonably representative of the whole cohort of patients, although BCR-ABL1-positive patients were slightly over-represented (Online Supplementary Table S4).
Deletions were more frequent than gains in all cytogenetic subgroups apart from high hyperdiploidy. Following the exclusion of probable constitutional copy number vari- ations, as described in the Online Supplementary Methods, a median of seven deletions (range, 0-52) and one gain (range, 0-29) were seen per patient sample.
In the 68/78 patients without a primary ploidy shift (de- fined as HoTr and high hyperdiploidy), large deletions on 9p were the most prevalent arm-level CNA, seen in 22%
T. Creasey et al.
(15/68) of cases (Figure 2, Online Supplementary Table S5). An additional copy of the Philadelphia chromosome was present in 12% (8/68) of patients (26% of BCR-ABL1-posi- tive cases) and 1q gains and monosomy 7 were each pres- ent in 10% (7/68) of samples.
Of the CNA in known driver genes, IKZF1 deletions were the most frequent abnormality, present in 51% (40/78) of cases. These were focal intragenic deletions in 19 cases, most commonly involving exons 4-7 (n=11) or exons 2-7 (n=4). Rarer IKZF1 deletions involved exons 4-8 (n=2), exons 2-8 (n=1) and one patient had biallelic IKZF1 loss in- volving exons 2-7 and 2-8. Focal IKZF1 deletions were al- most exclusively seen in patients with BCR-ABL1 (n=13) or CRLF2 rearrangements (n=5) (Online Supplementary Table S6). In the remaining cases, IKZF1 loss resulted from monosomy 7 (n=16) or del(7p) (n=5) (Figure 2, Online Sup- plementary Table S6).
The pattern of gene deletions varied by BCR-ABL1 status with a higher frequency of IKZF1 deletion in BCR-ABL1- positive ALL, as previously described,24 and a higher fre- quency of ETV6 and RB1 deletions in BCR-ABL1-negative
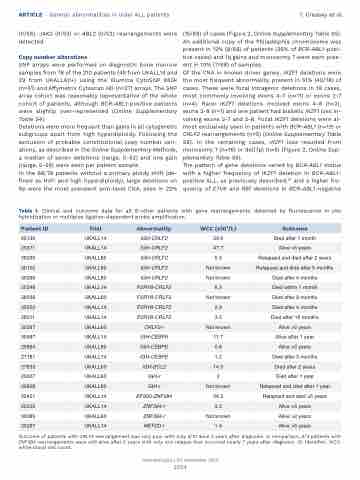
Table 1. Clinical and outcome data for all B-other patients with gene rearrangements detected by fluorescence in situ hybridization or multiplex ligation-dependent probe amplification.
Patient ID
Trial
Abnormality
WCC (x109/L)
Outcome
25130
UKALL14
IGH-CRLF2
33.6
Died after 1 month
25371
UKALL14
IGH-CRLF2
47.7
Alive >5 years
28235
UKALL60
IGH-CRLF2
5.3
Relapsed and died after 2 years
30102
UKALL60
IGH-CRLF2
Not known
Relapsed and died after 5 months
30299
UKALL60
IGH-CRLF2
Not known
Died after 4 months
25246
UKALL14
P2RY8-CRLF2
6.3
Died within 1 month
28039
UKALL60
P2RY8-CRLF2
Not known
Died after 9 months
25552
UKALL14
P2RY8-CRLF2
2.9
Died after 4 months
28011
UKALL14
P2RY8-CRLF2
3.5
Died after 16 months
30297
UKALL60
CRLF2-r
Not known
Alive >2 years
30487
UKALL14
IGH-CEBPA
11.7
Alive after 1 year
25894
UKALL60
IGH-CEBPD
0.8
Alive >5 years
27181
UKALL14
IGH-CEBPE
1.2
Died after 3 months
27833
UKALL60
IGH-BCL2
14.5
Died after 2 years
25907
UKALL60
IGH-r
2
Died after 1 year
29808
UKALL60
IGH-r
Not known
Relapsed and died after 1 year
25451
UKALL14
EP300-ZNF384
34.2
Relapsed and died >5 years
25235
UKALL14
ZNF384-r
3.5
Alive >5 years
30085
UKALL60
ZNF384-r
Not known
Alive >2 years
25267
UKALL14
MEF2D-r
1.4
Alive >5 years
Outcome of patients with CRLF2 rearrangement was very poor with only 2/10 alive 2 years after diagnosis. In comparison, 2/3 patients with ZNF384 rearrangements were still alive after 5 years with only one relapse that occurred nearly 7 years after diagnosis. ID: identifier; WCC: white blood cell count.
Haematologica | 107 September 2022
2054